Home



PROGRAM SUMMARY:
The WE-SPARK Cancer Program is an assembly of local researchers, healthcare professionals and community partners working together to build teams that strengthen our local cancer research programs and bridge collaborations with our neighbouring cancer centres. As the flagship program for WE-SPARK, we offer a number of cancer-related resources are found throughout the site that might be of interest. The Cancer Resource Hub provides the most credible and up-to-date links for researchers and our community. Our Research Registry enables community participation in ongoing research projects. Check out the page for current studies and eligibility. Additionally, visit our Cancer Research News section to stay up to date on local cancer research progress. Join our team here.
Mission: To establish Windsor-Essex as an international hub of cancer research excellence.
Vision: A united community bridging cutting edge research with world-class cancer care.
“Today, cancer patients live longer because of advancements stemming from research. Research outcomes have a global benefit, but there are additional impacts felt when research takes place locally. Research brings cutting-edge infrastructure to the community, delivers the latest innovative treatments to local patients, elevates knowledge of all healthcare providers, and attracts and retains the brightest professionals and students. Data shows that patients treated in institutions with active research programs have improved outcomes. Research saves lives. WCRG is committed to growing and strengthening our local cancer research programs and clinical trials to improve health outcomes for all those living in Windsor-Essex. We also have many talented professionals and students with brilliant ideas; and we lie at the nexus of US and Canada, making us unique in our ability to actively collaborate with our US partners. We are excited to be the flagship program under the WE-SPARK Health Institute and we look forward to continuing to build innovative research and health care excellence right here in our community.”
 | Caroline Hamm, M.D. FRCP(C) |
| Dora Cavallo-Medved, Ph.D. |
Caroline Hamm | Krista Naccarato | Laurie Freeman | |||
Dora Cavallo-Medved | Karen Metcalfe | John Trant | |||
Lisa Porter | Ken Schneider | Indryas Woldie |
Since 2012, the WCRG has been the spark in igniting and elevating cancer research in Windsor-Essex through collaboration, engagement and community outreach. Here is a summary of WCRG metrics from 2020.
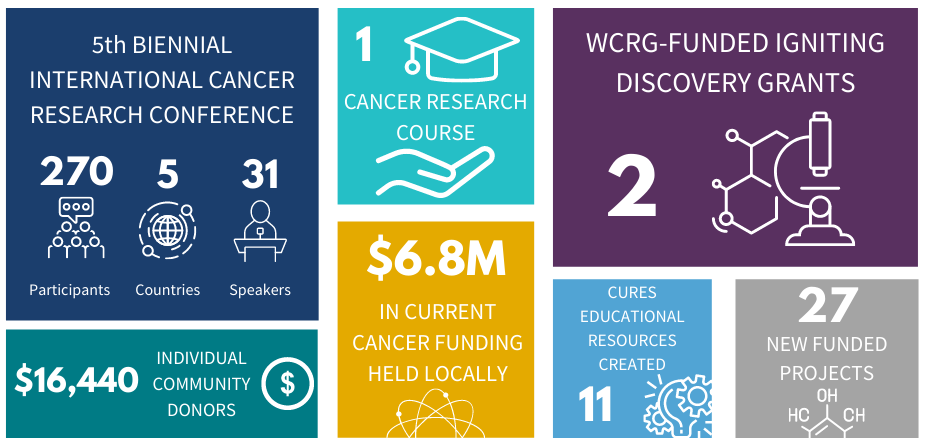
The 2020-2021 Cancer Research Impact can be found in WE-SPARK's latest Impact Report. View WCRG's past impact reports here.
Our cancer research program is made up of local researchers, clinicians, healthcare providers, research assistants and associates, administrative staff, and student trainees, across a range of disciplines. Additionally, our program has represenetation from all of WE-SPARK's partners - University of Windsor, St. Clair College, Hôtel-Dieu Grace Healthcare, and Windsor Regional Hosptial. Each of our members plays a key role in the cancer research network, bringing to the table different areas of expertise, experiences, connections, and ideas.
.png)
 | WCRG launched our Think Tank: Cancer series in 2016, which have since extended to all areas of health with the launch of WE-SPARK in 2019. Learn more about our Think Tanks here. |
 | In our impact reports, you will find a high level overview of our yearly successes. Check them out: 2012-2015, 2016, 2017, 2018. |
| Our Ambassadors of Hope program was launched to recognize members of our cancer research ecosystem that demonstrate continuous committment to moving cancer research forward in Windsor-Essex. With the launch of WE-SPARK, this program will recognize key contributors across all areas of health. |
 | In 2012, WCRG hosted it's first Biennial International Cancer Research Conference that became a bi-yearly event. Check out our conference page for more information. |
| In collaboration with Snapd Windsor, now known as Neighbur, many of our researchers completed a Q&A that allowed the community to get to know a bit about them and their research. Check them out! |
If you're interested in volunteering with WE-SPARK, check out our volunteer network webpage!
If you're looking to volunteer with our cancer research program, we encourage you take various courses in science communication, research ethics, principals of research, etc. Here are a few examples:
| Institution | Course | Description |
University of Windsor | SCIE 3750: Cancer Undergraduate Research Education (CURES) | This course engages students in learning about the science of cancer and directions in cancer research. Students will interact with scientists conducting a spectrum of multidisciplinary research and will work in collaborative teams to design and implement tools to communicate cancer research to the public, to patients, to other students, and to the government. These ideas will help move cancer research forward in Windsor Essex. |
Our cancer research program prioritizes community outreach as an essential component to our success. Throughout the last decade, we have participated in a variety of such events (see list below). NOTE: Due to the COVID-19 restrictions, many of these events have been paused or converted to a virtual format. Check out our social media pages (Facebook, Twitter) and our events section for updates!
To see more photos, check out our photo gallary.


















Today, cancer patients live longer because of advancements stemming from research. WCRG is excited to build and strengthen our current local cancer research program to better serve the patients of the Lake Erie-St. Clair region. And we are dedicated to making a difference in the fight against cancer. Donations made to the Windsor Cancer Research Group are designated to our Grants Program. With your help, these grants will help grow our research community, ignite new discoveries, address gaps in healthcare needs, generate and disseminate knowledge, spark collaborations and improve the health of our community. Thank you to all our supporters, we couldn't do it without you!
 | Research drives discoveries and provides solutions. Research supports health and wellness. Research trains the next generation. Research betters clinical practice. Research fuels the economy. Research saves lives. |
 Believer |  Friend |  Champion |  Visionary |
Marcella Beneteau Michael J. Dufresne Jo Anne Lambing Wonder Broads of Windsor-Essex | Lawerence Delmore Peter Dunn Heather Metcalfe | Gang Lu | Windsor Cancer Research Windsor Cancer Centre |
Past Supporters
|
Charitable Tax receipts for gifts $10 and over will be issued by mail.Reg.#10816 2611 RR0001